|
Research

Epigenome maintenance during cell proliferation and in response to stress
DNA replication is a central event of cell division and is linked to cell cycle regulation and the cellular stress response in many ways. The precise and coordinated duplication of genetic information is critical for genome stability.
We study the coordination and dynamics of the factors involved in the replication and repair of the genome using multidimensional time-lapse microscopy in combination with photobleaching/activation techniques. We elucidated fundamental changes in the progression of genome replication in pluripotent embryonic stem cells versus differentiated cells. Altogether, our data argue against the existence of a stable replisome complex in cells and uncover the dynamic nature of the replisome components. We proposed that genome replication progresses by activation of neighboring replication origins by a “domino effect” involving local changes in chromatin accessibility. Using new methods for targeted manipulation of chromatin (com)position, we found that the level of histone acetylation and the nuclear position control the replication timing of constitutive and facultative heterochromatin. Using superresolution microscopy, we were able to resolve and quantify individual replicons in single somatic cells as well as embryonic stem cells in 3D and developed a minimal comprehensive model for DNA replication in single human cells.
In parallel, we have been investigating the links between DNA replication and damage repair, including the recruitment of DNA synthesis factors to DNA damage sites. Using genomic approaches, we found that genetic and epigenetic features regulate damage susceptibility and repair kinetics. Using superresolution microscopy, we found that 3D spatial clustering of DNA damage (nano)foci by CTCF is required for repair.
Epigenome reprogramming during normal cellular differentiation and in disease
Most cells of multicellular organisms contain identical genetic information but differ in their epigenetic information, including DNA base modifications (in mammals, 5-methylcytosine, 5-hydroxymethylcytosine, formyl-cytosine and carboxy-cytosine) and histone modifications. Our focus is on the role and regulation of DNA modifications.
Methyl-cytosine binding proteins (MeCP2, MBD1-4) recognize and bind to sites of DNA methylation and recruit other chromatin modifiers. We found that MBD binding to methylated cytosines protects the modification from being further modified by the TET dioxygenases and, thus, MBDs act as “guardians of the epigenome”. This protection is disrupted in MeCP2 deficient animals as models for Rett syndrome and leads to increase transcriptional noise from tandem and interspersed DNA repeat elements with increased retrotransposition. We have found that MBDs induce large-scale heterochromatin reorganization during differentiation and are investigating the underlying mechanism using a combination of in vitro and in vivo approaches in embryonic stem cells, induced pluripotent stem cells and differentiated cells. We also found that MeCP2 is posttranslational modified in brain by PARP-1 and this decreases its chromatin binding.
Targeting and tracing cellular structures and functions in living cells
We have studied and optimized cell penetrating peptides (CPPs) to visualize and manipulate cellular functions and uncovered the fundamental mechanism of non-endocytic crossing of the plasma membrane by CPPs.
In parallel, we have shown that alpaca-derived nanobodies fused to fluorescent proteins, so called chromobodies, can be expressed in mammalian cells/organisms and used to target and trace antigens in living cells and are very useful to study protein interactions in living cells and throughout the cell cycle.
We combined CPPs with nanobodies to create cell permeable nanobodies allowing targeted immunolabeling and antigen manipulation in living cells.
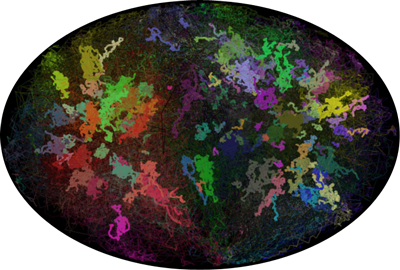
The interactive 3D viewer displays the chromatin structure (76 chromosomes, HeLa cells) resulting from a modified random loop model with about 104,000 beads.
Check it out: http://sim.bio.tu-darmstadt.de/sim/viewer_3d/dna/
We are members of:
Our group is funded by:

|

